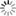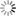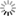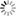Import
Importing your Images into OME
OME allows users to import multi-dimensional microscope images into the OME data hierarchy (images --> datasets --> projects). These images are available for further examination and processing. We currently support Applied Precision SoftWorx (Deltavision), Metamorph STK and HTD, Molecular Dynamics GEL, Zeiss LSM, Bio-Rad PIC, DICOM (medical image format), TIFF, and OME-XML image formats. As a result of our partnership with the developers of VisBio their viewer can, since version 3.00, import all of the many common image formats it understands directly into OME. They have also developed a similar plugin for ImageJ.
Many image formats record information that is known at acquisition time about the biology work-flow, e.g. a detailed description of the microscope, what filters were used for each channel, etc. During image import, this information is stored with the image's pixels as meta-data, and is accessible through OME's various interfaces.
Importing datasets with many large images can take a long time. You can view the progress of your import process in the Tasks page.